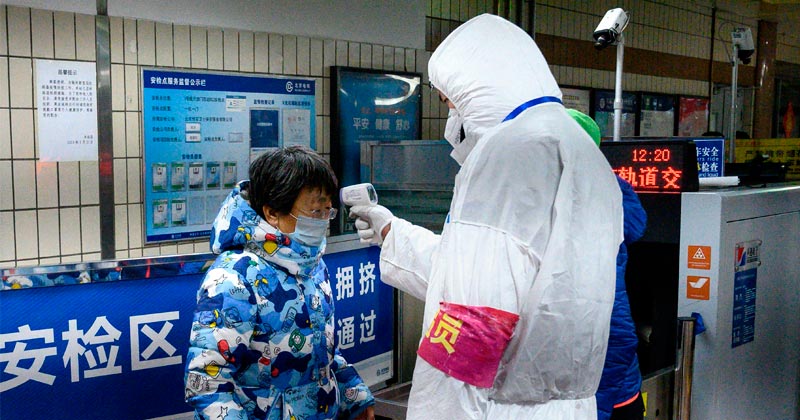
IMAGE CREDITS: NOEL CELIS/AFP VIA GETTY IMAGES.
CORONAVIRUS CONTAINS “HIV INSERTIONS”, STOKING FEARS OVER ARTIFICIALLY CREATED BIOWEAPON
‘The virus even responds to treatment by HIV medications’
Over the past few days, the mainstream press has vigorously pushed back against a theory about the origins of the coronavirus that has now infected as many as 70,000+ people in Wuhan alone (depending on whom you believe). The theory is that China obtained the coronavirus via a Canadian research program, and started molding it into a bioweapon at the Institute of Virology in Wuhan. Politifact pointed the finger at Zero Hedge, in particular, though the story was widely shared across independent-leaning media.
The theory is that the virus, which was developed by infectious disease experts to function as a bio-weapon, originated in the Wuhan-based lab of Dr. Peng Zhou, China’s preeminent researcher of bat immune systems, specifically in how their immune systems adapt to the presence of viruses like coronavirus and other destructive viruses. Somehow, the virus escaped from the lab, and the Hunan fish market where the virus supposedly originated is merely a ruse.
Now, a respected epidemiologist who recently caught flack for claiming in a twitter threat that the virus appeared to be much more contagious than initially believed is pointing out irregularities in the virus’s genome that suggests it might have been genetically engineered for the purposes of a weapon, and not just any weapon but the deadliest one of all.
In “Uncanny similarity of unique inserts in the 2019-nCoV spike protein to HIV-1 gp120 and Gag“, Indian researchers are baffled by segments of the virus’s RNA that have no relation to other coronaviruses like SARS, and instead appear to be closer to HIV. The virus even responds to treatment by HIV medications.
For those pressed for time, here are the key findings from the paper, which first focuses on the unique nature of 2019-nCoV, and then observe four amino acid sequences in the Wuhan Coronavirus which are homologous to amino acid sequences in HIV1:
Our phylogentic tree of full-length coronaviruses suggests that 2019-nCoV is closely related to SARS CoV [Fig1].
In addition, other recent studies have linked the 2019-nCoV to SARS CoV. We therefore compared the spike glycoprotein sequences of the 2019-nCoV to that of the SARS CoV (NCBI Accession number: AY390556.1). On careful examination of the sequence alignment we found that the 2019- nCoV spike glycoprotein contains 4 insertions [Fig.2]. To further investigate if these inserts are present in any other corona virus, we performed a multiple sequence alignment of the spike glycoprotein amino acid sequences of all available coronaviruses (n=55) [refer Table S.File1] in NCBI refseq (ncbi.nlm.nih.gov) this includes one sequence of 2019-nCoV[Fig.S1]. We found that these 4 insertions [inserts 1, 2, 3 and 4] are unique to 2019-nCoV and are not present in other coronaviruses analyzed. Another group from China had documented three insertions comparing fewer spike glycoprotein sequences of coronaviruses . Another group from China had documented three insertions comparing fewer spike glycoprotein sequences of coronaviruses (Zhou et al., 2020).
We then translated the aligned genome and found that these inserts are present in all Wuhan 2019-nCoV viruses except the 2019-nCoV virus of Bat as a host [Fig.S4]. Intrigued by the 4 highly conserved inserts unique to 2019-nCoV we wanted to understand their origin. For this purpose, we used the 2019-nCoV local alignment with each insert as query against all virus genomes and considered hits with 100% sequence coverage. Surprisingly, each of the four inserts aligned with short segments of the Human immunodeficiency Virus-1 (HIV-1) proteins. The amino acid positions of the inserts in 2019-nCoV and the corresponding residues in HIV-1 gp120 and HIV-1 Gag are shown in Table 1.
The first 3 inserts (insert 1,2 and 3) aligned to short segments of amino acid residues in HIV-1 gp120. The insert 4 aligned to HIV-1 Gag. The insert 1 (6 amino acid residues) and insert 2 (6 amino acid residues) in the spike glycoprotein of 2019-nCoV are 100% identical to the residues mapped to HIV-1 gp120. The insert 3 (12 amino acid residues) in 2019- nCoV maps to HIV-1 gp120 with gaps [see Table 1]. The insert 4 (8 amino acid residues) maps to HIV-1 Gag with gaps.
Why do the authors think the virus may be man-made? Because when looking at the above insertions which are not present in any of the closest coronavirus families, “it is quite unlikely for a virus to have acquired such unique insertions naturally in a short duration of time.” Instead, they can be found in cell identification and membrane binding proteins located in the HIV genome.
Since the S protein of 2019-nCoV shares closest ancestry with SARS GZ02, the sequence coding for spike proteins of these two viruses were compared using MultiAlin software. We found four new insertions in the protein of 2019-nCoV- “GTNGTKR” (IS1), “HKNNKS” (IS2), “GDSSSG” (IS3) and “QTNSPRRA” (IS4) (Figure 2). To our surprise, these sequence insertions were not only absent in S protein of SARS but were also not observed in any other member of the Coronaviridae family (Supplementary figure). This is startling as it is quite unlikely for a virus to have acquired such unique insertions naturally in a short duration of time.
The insertions were observed to be present in all the genomic sequences of 2019-nCoV virus available from the recent clinical isolates. To know the source of these insertions in 2019-nCoV a local alignment was done with BLASTp using these insertions as query with all virus genome. Unexpectedly, all the insertions got aligned with Human immunodeficiency Virus-1 (HIV-1). Further analysis revealed that aligned sequences of HIV-1 with 2019-nCoV were derived from surface glycoprotein gp120 (amino acid sequence positions: 404-409, 462-467, 136-150) and from Gag protein (366-384 amino acid) (Table 1). Gag protein of HIV is involved in host membrane binding, packaging of the virus and for the formation of virus-like particles. Gp120 plays crucial role in recognizing the host cell by binding to the primary receptor CD4.This binding induces structural rearrangements in GP120, creating a high affinity binding site for a chemokine co-receptor like CXCR4 and/or CCR5.
A good recap of the findings was provided by Dr. Feigl-Ding, who started his explanatory thread by pointing out that the transmission rate outside China has surpassed the rate inside China.
READ THE REST HERE >> https://www.infowars.com/coronavirus-contains-hiv-insertions-stoking-fears-over-artificially-created-bioweapon/